Systems thinking
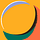
If I were given the challenge to design the curriculum for an introductory biology course, I would include a couple of lectures on Systems Thinking. This would be less abstract than the biologist Ludwig Von Bertalanffy’s General Systems Theory of the late 1960s -- it would encapsulate “models, principles, and laws that apply to generalized systems or their subclasses, irrespective of their particular kind, the nature of their component elements, and the relationships or “forces” between them.” Nor is it synonymous with what is recognized as systems biology by molecular biologists – the generation of large transcriptomic datasets and their subsequent analysis to identify patterns or networks. O’Malley and Dupre (2005) described this as ‘pragmatic’ systems biology. Mine would be their ‘systems-theoretic’ biology with an intent to identify principles that are broadly applicable to different specific biological systems and (perhaps) at multiple hierarchal levels. As I started to think about this, I realized it dovetails very well with Craver’s depiction of a mechanism underlying a phenomenon, that I discussed in this post.
If you recall, Craver and Darden proposed that experimentalists devise multi-level schema of how system components interact with each other to better explain how a phenomenon is caused. That is systems thinking in a nutshell. It is both art and science – making predictions about systems behavior by developing an increasingly deep understanding of underlying structure.
My intention for adding this to a biology curriculum would be to expose students to a new/different way of thinking about biology. Not to stop at a collection of trees (e.g., regulation of tryptophan biosynthesis in a cell) but think about the network that regulates the rates of biosynthesis of all 20 amino acids such that each are in balance with each other and with the cell’s demands for protein synthesis.
What are some fundamentals?
Vantage point is one essential fundamental. Per Craver and Darden’s concept of schema consisting of a nested hierarchy, systems thinking requires detailed analyses of sets of mechanisms at multiple levels to understand overall system dynamics
Some important principles include:
It is the set of interacting components in a system (and their responses to internal or external perturbations) that determines system behavior, rather than external forces per se.
Closed feedback loops are a fundamental structure regulating system behavior.
Many subsystems can be visualized as a series of stocks/pools that have levels/concentrations with flows both into and out of them. It is the infrastructure of stocks and flows that provide the basis for feedback loops that control the system.
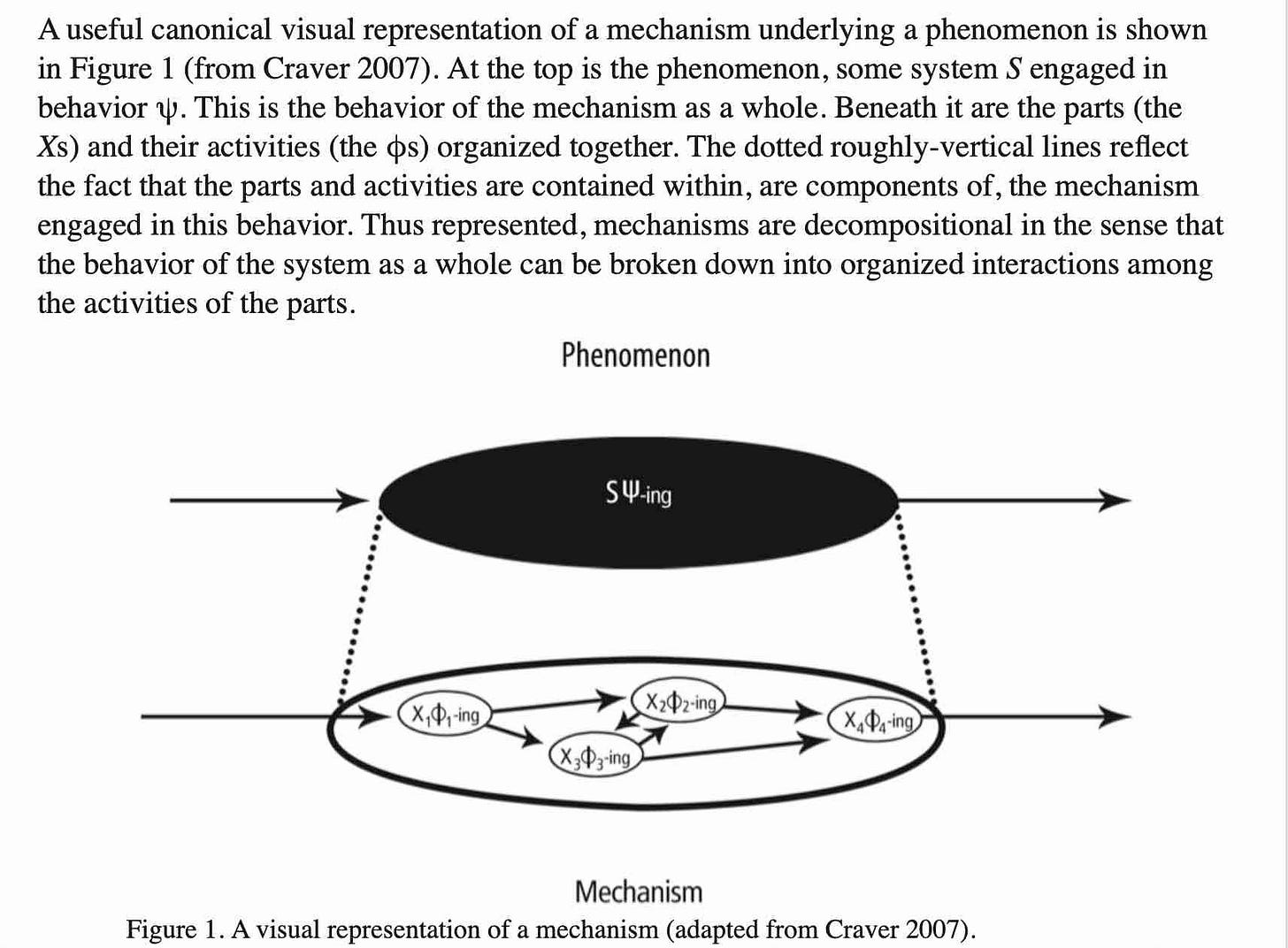
SIR epidemiological models (with stocks of Susceptible, Infected, and Resistant individuals) were first developed 100 years ago by Kermak and McKendrick (and remained a mainstay of epidemiological modeling at the inception of the COVID-19 pandemic!). Here is a graphical example of a stock & flow SIR model developed in 2020 using the simulation software Vensim.
You can see the basic structure in the 5 black Stocks, with flows between them. The flows are calculated as simple algebraic expressions, using values from elements with arrows pointing to the flows. For example the flow Becoming Symptomatic is calculated as Infected asympto/Incubation period. The modeler sets a time step over which flows and stocks are updated, effectively integrating dynamics over time. Homer has added a number of other factors whose values can be adjusted and their effects on the basic system evaluated. [Note: epidemic modeling over past 5 years has moved significantly towards individual agent-based models that incorporate stochasticity and temporal & spatial heterogeneities]
Simulation modeling
Thinking about systems and their component interactions raises consciousness regarding all strata of biological organization. Taking the next step with modeling simulators as simple as Vensim can provide predictions of short- and long-term consequences of altering components upon the system’s dynamics. Interfacing such modeling with experimentation to test model predictions can deepen our understanding of mechanisms.
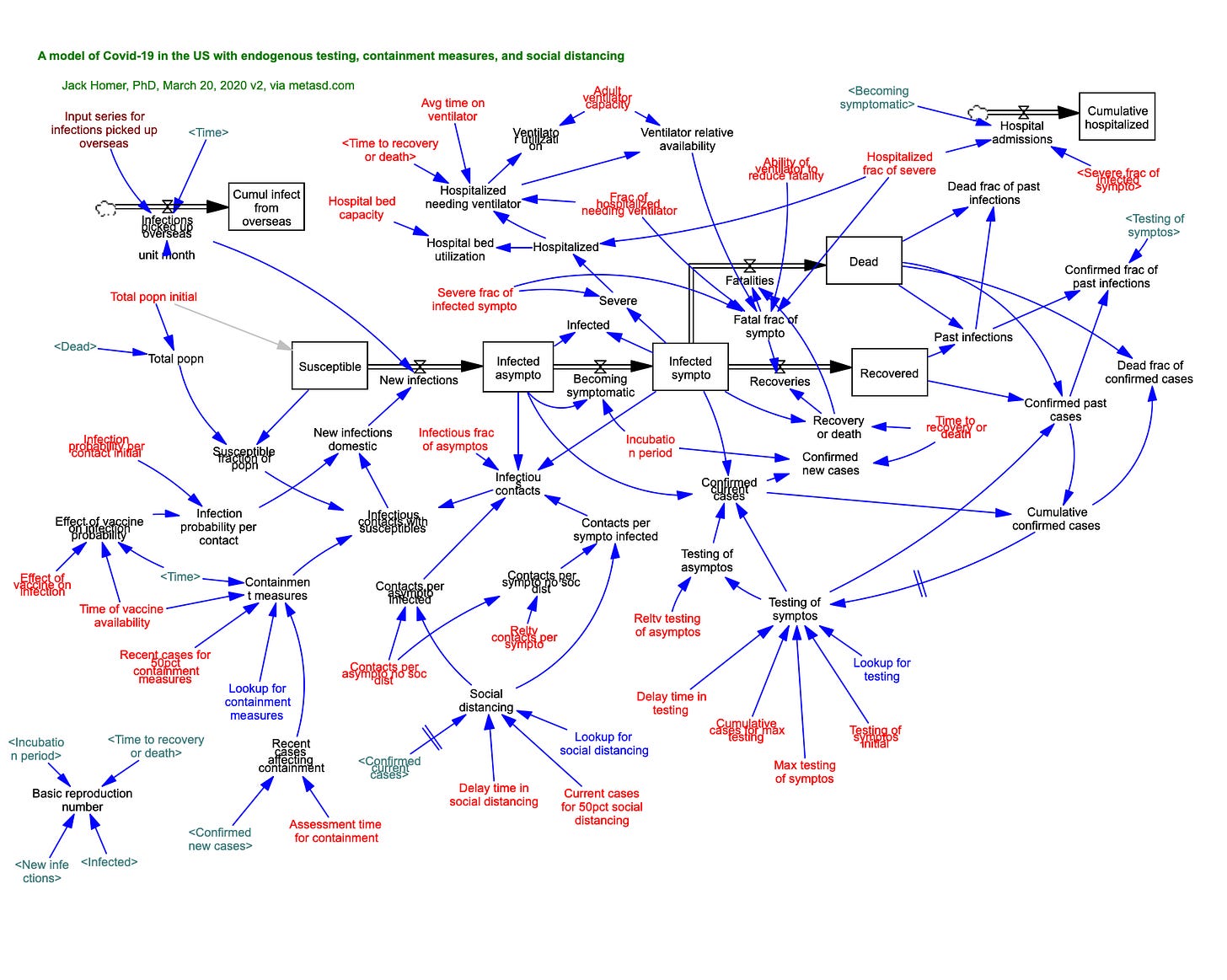
As one example, I constructed a simple model of this type to compare the fitness of a bacterium that shifted to a dormant state under nutrient starvation to one that did not. I varied the time interval between nutrient pulses and did sensitivity analyses in which the organisms’ kinetic parameters were systematically changed under the hypothesis that longer intervals would favor dormancy. The conclusion was that the selection coefficient for dormancy was most sensitive to the time constant for conversion of dormant cells back to actively growing ones, and the magnitude of the death rate of cells which did not undergo the starvation response. Subsequently, a Master’s student, Militza Carrero-Colón tested these predictions. Sadly, they were not supported – but it does represent a case of how even simple simulation modeling can inform subsequent experimentation
I also used simple graphically oriented simulation models such as the SIR models in teaching some of my microbiology courses at Purdue. For purely undergraduate courses, I built the model in the background and presented the students with a dashboard whereby they could control the simulation parameters.
For advanced undergraduate/graduate student course, I gave students a primer on how to construct models in software like Vensim and had them do a class project of their own design.
I found that engaging students with dynamic modeling of this type had some important outcomes:
They could experiment/play with the models. They could ask ‘what-if’ questions by changing paramaters and observing how system behavior changes.
A simulation model functions as a thought-organizing device (per Craver & Darden’s schema). For students, the value arises from a deeper understanding of how variables impact system behavior that static lectures to them cannot deliver. For researchers, the detailed model/schema may demand assembly of experts from other disciplines who can delve into specific subsystem assemblies. It is a great means to foster interdisciplinary collaboration as members gain knowledge from each other at the disciplinary boundaries of the system.
On feedback – negative vs positive
Most feedback loops in systems are negative and act to modulate rates and/or dampen oscillations. However, there are examples of positive feedback. Bacterial growth is an obvious one – it is auto-catalytic. The product of growth is more units that can catalyze growth. Of course, in a real-world system the depletion of an essential growth nutrient is a negative feedback that slows or stops growth rate.
A concern in Earth climate science is the fate of organic C sequestered in permafrost at high northern latitudes. The amount (~1500 Gt) is approximately 40 times the amount of fossil fuels burned each year. The rise in atmospheric CO2over the last 200 years has led to temperature increases at these high latitudes that have begun to melt permafrost, thereby making the organic C available for microbial metabolism. Current estimates are that ~10% of permafrost has thawed. If anoxic, then methane (another greenhouse gas) is produced, but once oxic, organic C will be mineralized to more CO2. This will represent a positive feed-forward in which thawing of permafrost leads to more CO2 release to the atmosphere which will drive an accelerated rate of warming and permafrost thawing.
Complex systems
Once one embraces systems thinking, the concept of complex system emerges [pun intended]. In 2009, I spent three days in a Washington DC hotel at a workshop for the U.S. Department of Energy’s Office of Biological and Environmental Research on complex systems science. I was looking forward to this as I had been engaged in systems thinking for a decade or so. However, it felt like significant time each day was spent re-litigating what made a system ‘complex’ rather than merely ‘complicated.’
Complex systems are thought to have ‘emergent’ properties – the self-organized formation of collective structures and behaviors that are distinct from those of individual entities that comprise the system. In microbial ecology some examples of emergent properties are:
Biofilm Formation / Spatial Self‑Organization
Metabolic Cooperation (Cross‑Feeding)
Community Resilience to environmental perturbations
More generally in macrobe ecology, these types of phenomena are considered emergent properties:
1. Tipping points that trigger large regime shifts in the system
2. Trophic cascades, where the effects of a predator ripple down through a food web and may even impact the physical landscape.
3. Functional Redundancy and Response Diversity
4. Biogeochemical cycles of nutrients and energy
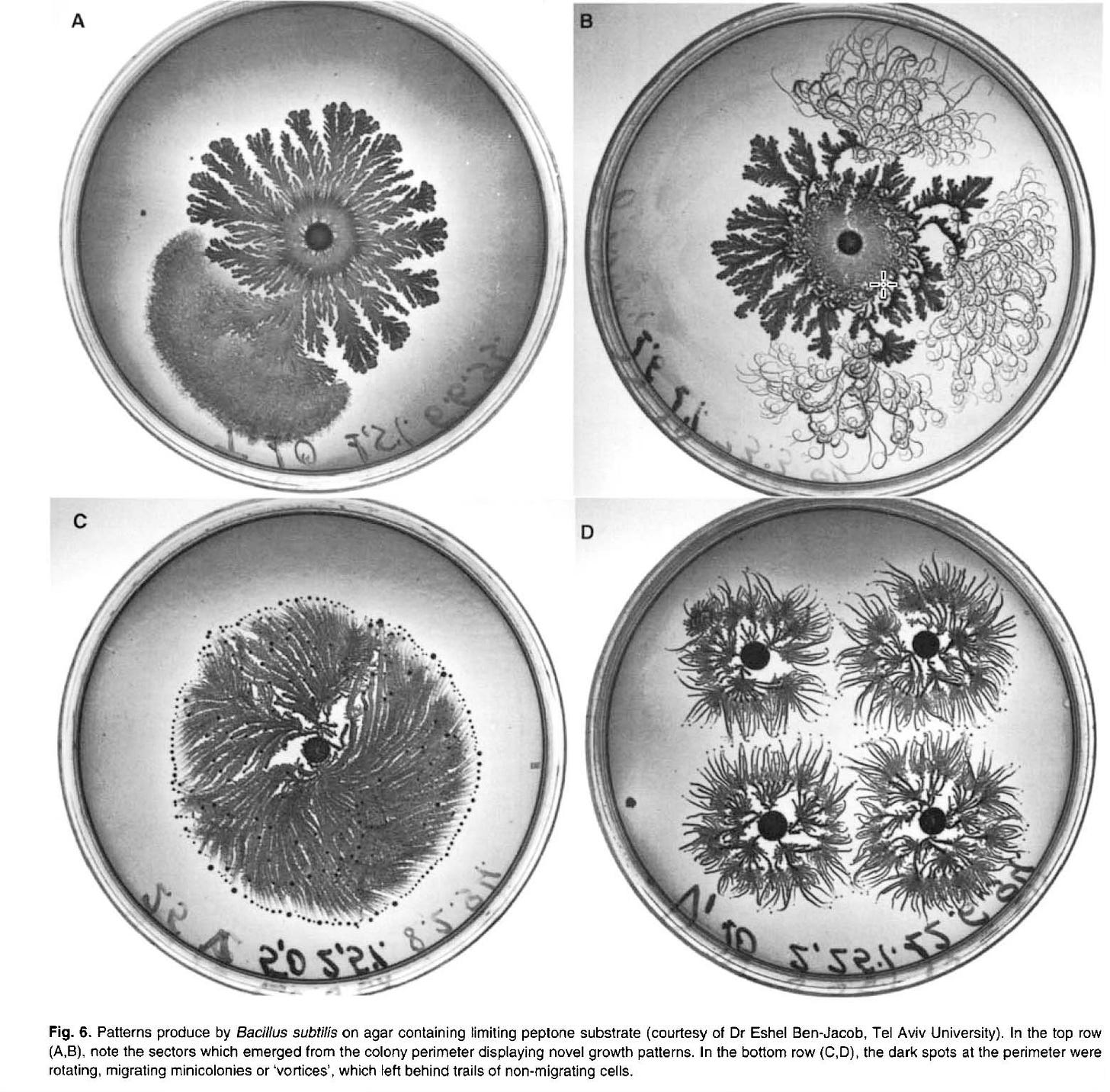
Defining or identifying what constitutes an emergent property can be fraught and contentious. In my view, a system is ‘complex’ if the relationships between system constituents are not strictly additive or linear. Nonlinear dynamics can result in even small perturbations producing a very large effect on system behavior and can lead to chaos (in a mathematical sense), periodic or nonperiodic oscillatory behavior, or the type of fractal organization seen in bacterial colonies.
Earth System Science
Lovelock and Margulis (1974) created quite a stir 50 years ago: “The notion of the biosphere as an active adaptive control system able to maintain the Earth in homeostasis we are calling the ‘Gaia’ hypothesis.” I think I was turned off by the mystical connotations at the time as well as the idea of Earth as a living organism. If we just call it Earth Systems Science and emphasize the physical, chemical and biological feedback loops that regulate processes I am OK with it. But there is incredible complexity that we very incompletely understand. Consider that despite the advances in physics models, satellite sensing coverage and computational power made in the last 50 years, predicting the weather 10 days from now is no better than a coin flip.
I came across a book written by the eminent microbial molecular and evolutionary biologist, W Ford Doolittle, Darwinizing Gaia. It is open access and I highly recommend it.. He includes very nice chapters that summarize the Gaia Hypothesis and the Modern (1940s) Synthesis of Evolution. Margulis extended Lovelock’s original Gaia hypothesis from a homeostatic mechanism to one that is autopoietic and evolutionary. Doolittle was a vocal Gaia naysayer – on the basis that Earth can’t be an ‘organism,’ because natural selection operates when organisms reproduce and Earth does not. However, in this book he is at least open to arguments for Gaia on the basis that selection *might* also operate not only via reproduction but also by persistence.
Summing up
I remain enthusiastic regarding all biologists engaging in systems thinking, developing schema of how they perceive the mechanisms that underlie their systems and even engaging in systems modeling of stocks and flows to improve their intuition about their dynamics. However, I also recognize that deeper steps require much more and if I look at the literature, progress seems rather slow. One impediment I articulated in an earlier post on Mathematical Modeling:
“Progress requires effective intellectual engagement between microbial ecologists and mathematicians. It would be rare for a single individual to function at the highest level in both ecology and mathematics.”
The nature of complex systems (i.e., those that contain emergent properties) is that they can exhibit nonlinear, unstable and aperiodic dynamics. Investigators may attribute time series with such properties as “noise” or measurement error, biased by the long-standing idea that populations and communities exhibit a “balance of nature.” But what if the actual dynamics in systems are chaotic, with instabilities and sensitivity to initial conditions? That requires a different set of mathematics with which even mathematically-inclined biologists are uncomfortable.
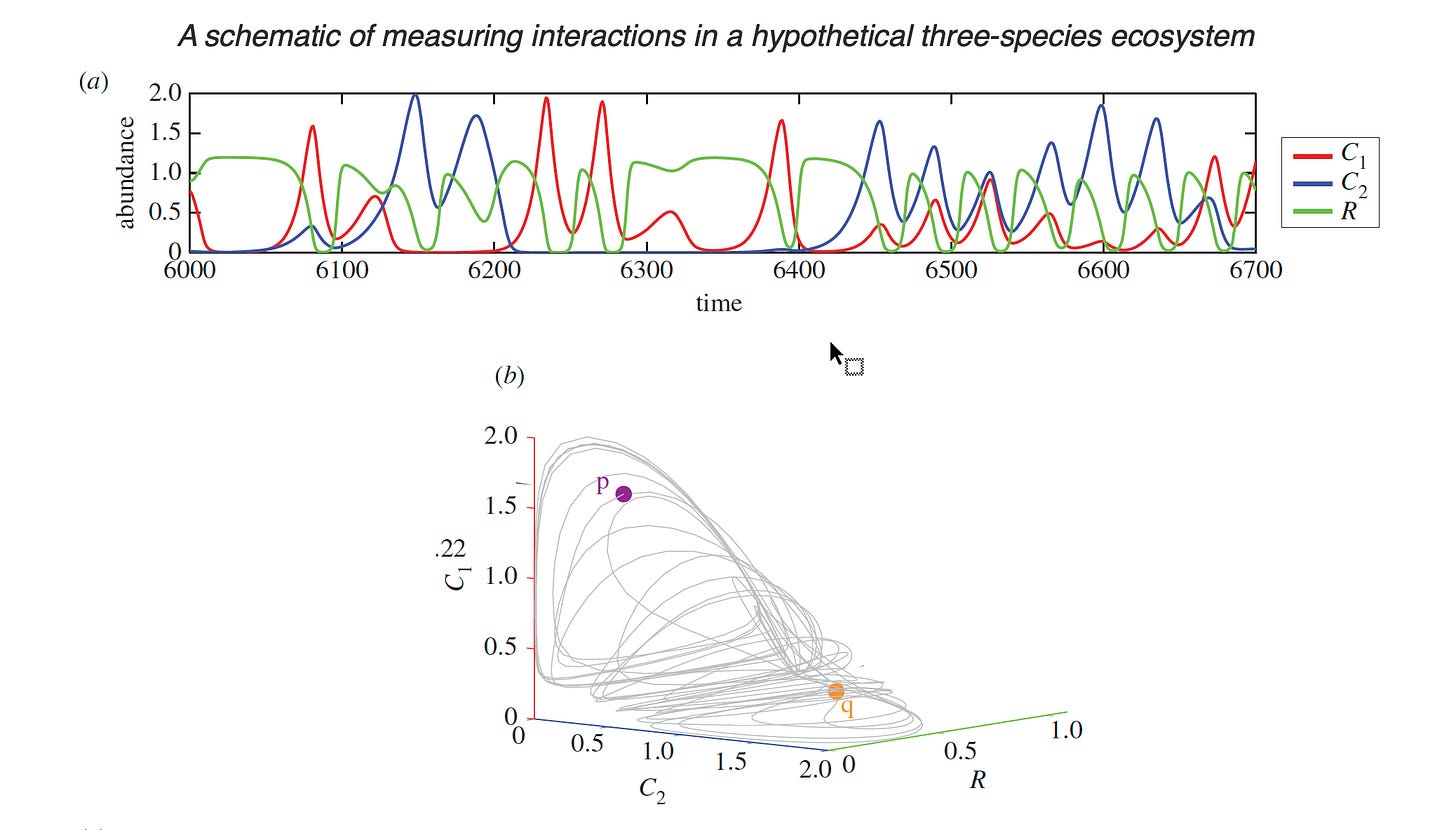
I have found George Sugihara’s work most interesting in this regard. Sugihara trained under the late, great Robert May and starting in 1990 investigated nonparametric mathematical methods of exploring time-series data to reconstruct ‘attractors’ that disentangle apparent ‘chaos.’ This approach most recently has been called Empirical Dynamic Modeling. In the end, this approach illuminates phenomena but the attractors may point to high-level mechanisms such as competition or predation.
Source: Tracking and forecasting ecosystem interactions in real time
Today’s Moment of Zen
We are leaving home in about a week for a two week cruise on the Danube River, from Romania up to southern Germany. This got me thinking about earlier trips to Eastern Europe, and I pulled this from my photo archive, taken on a one week road trip through Slovakia prior to the 2006 ISME meeting in Vienna.