Mathematical Modeling in Microbial Ecology
A little history. As a high-school student I loved mathematics. I think this was due in large part to the enthusiasm and rigor of my teacher. However, a couple of semesters at university where I was taught by graduate teaching assistants wrung all of that love out of me and I did not progress beyond calculus. I regret that, as I later saw how facility in mathematics could have improved my science. About 30 years ago, I read the esteemed limnoecologist WT Edmondson’s reflections “The Uses of Ecology: Lake Washington and Beyond.” As a high school student working in GE Hutchinson’s lab at Yale prior to entering college, he asked Hutchinson for advice on courses. Hutchinson’s answer was “keep taking math courses until you fail one.” Sage advice – mathematics is a tool that you should develop to the limit of your abilities. (Although I imagine the pre-med students I advised at Purdue would not have been swayed by this argument).
This post directly follows my thoughts on theorizing. A theory need not be mathematical, but if it is expressed in mathematical language, it has the virtue of generating quantitative predictions which can then be empirically tested (that is, Popper’s method of advancing science). Some mathematical models may be solved analytically, but more popular are simulation models in which a system of ordinary differential equations are solved via numerical integration.
History
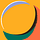
The ’classic’ models of the early days of ecology were the Lotka-Volterra predator-prey equations and the Verhulst (logistic) growth equation, both developed in the 1920s. There is a generalized form of Lotka-Volterra for any number of species that compete or otherwise interact with each other. This comprises a set of ordinary differential equations (one for each species) in which the maximum growth rate of the species is provided and there is an interaction matrix that describes the interactions between each pair of species. Whereas the predator-prey equation generates limit cycles, the more complex interaction matrices of the generalized equations may result in limit cycles, point attractors, or chaos.
Phase plane of Predator vs Prey in a Lotka Volterra model, illustrating limit cycles whose amplitude depend upon initial conditions of prey and predators (X0).
The logistic growth equation remains broadly used in ecology, although in my opinion the Monod equation
dX/dt = X *µmax * S/(K+S)
coupled with a specific accounting of limiting substrate consumption
dS/dt = -(µX/Y)
is far superior.
[X=biomass, S=substrate concentration, µ=instantaneous growth rate, K=concentration of S necessary for 0.5 µmax and Y=yield biomass per unit substrate]
The latter half of the 20th Century produced some important advances in theoretical ecology and modeling. Howard T. Odum was a driving force behind the development of “systems ecology” and simulation models of the energy flows through ecosystems. These began as electrical circuit diagrams (he used batteries, wires, resistors and capacitors as analogs of ecosystems in his earliest work!) but developed into a series of linked ‘reservoirs’ with regulated inflows and outflows. His work exemplified the top-down approach to ecology and his 1994 book, Ecological and General Systems, remains a useful compendium of the mathematical, kinetic, and energetic bases for the systems ecology approach.
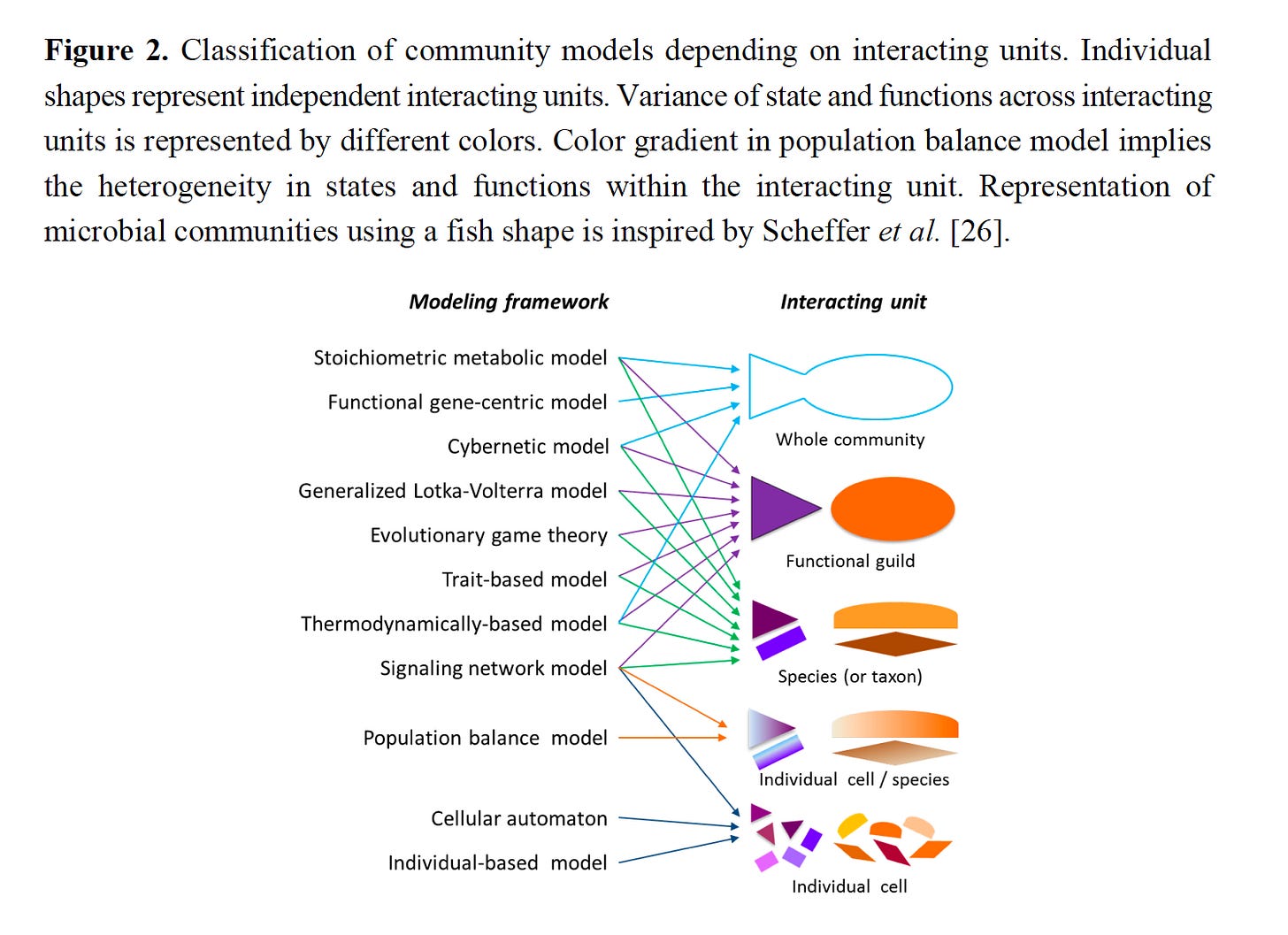
In my opinion, Robert May had an outsized influence on mathematical approaches to ecology. Trained as a theoretical physicist, the work of Hutchinson and Macarthur led him to apply his mathematical expertise to fundamental questions in ecology starting in the late 1960s. His 1973 monograph, Stability and Complexity in Model Ecosystems, explored the diversity-stability debate and demonstrated that stability was far from a given in randomly-assembled communities. This opened an entire field regarding what characteristics make systems robust and stable and what types of conditions will produce transitions in community composition. The conventional wisdom at the time was that shifts in community composition were driven by stochastic changes in exogenous factors. However, his analysis of simple sets of difference equations showed that ‘internal’ dynamics could of themselves generate highly complex community responses, venturing as far as what is now called chaos. The generality of this idea remains unsettled, although nonlinear dynamics continues to be an important basis for modeling the complex behaviors observed in natural communities. He and Sugihara (1990) later developed an approach to make predictions regarding the trajectories of nonlinear dynamic systems from time series data. Sugihara call this approach Empirical Dynamic Modeling.
Models in Microbial Ecology
The record is rather thin here. Of course, microbial ‘compartments’ are part of systems-level ecosystem models, although their representation is at a very coarse level. Biofilms are a natural for Individual-Based Modeling, given their importance in dental caries and a variety of engineered ecosystems. In this approach, each individual cell is modeled in space and assigned ‘rules’ that govern its behavior. This is intriguing as it can get to the microenvironment scale. However, these models can quickly become complex and hence computationally expensive unless the rules (and spatial scale) are quite simple. Jan-Ulrich Kreft and his colleagues have developed a simulator iDynoMics (Cockx et al., 2024) that can be downloaded from GitHUB.
Genome-scale metabolic flux models came into vogue as complete genome sequences of bacteria became readily available. These are based on flux balance analysis (FBA), developed by the chemical engineer Bernard Palsson and his colleagues. From the genome, a matrix of reactions is constructed. Constraints are imposed and a solution is obtained to maximize an objective function (for example, growth). The output is the flux of metabolites through the various metabolic pathways. This is a great approach for questions in chemical engineering reactors, where microbes may be in steady-state or balanced growth conditions. However, note that the FBA only produces a solution at steady state – that almost never occurs in natural microbial communities.
In my opinion, these metabolic flux models miss the mark by focusing on containing all the details (enzymes) of metabolism, many of which are common to all organisms. Going back to the mantra of Clements that ecology is the study of stimulus and response, I believe that the focus should be on the regulation of cell activities by the extant physico-chemical conditions. Based on the collaborations I had with Doraiswami Ramkrishna (Ramki), an eminent chemical engineer at Purdue, on modeling a microbe in a bioreactor, I came to believe that the key was to simulate the effects of regulation while minimizing the details. His approach is cybernetic modeling which implicitly represents regulatory mechanisms, and calculates their impact via cybernetic variables that constrain metabolic processes to achieve an (assumed) goal (for example, maximize growth rate or minimize death rate). Of particular importance in environments containing limiting or transiently-available concentrations of heterogeneous nutrient resources would be simulating regulation of different transporters. Ramki’s cybernetic models contained ‘enzymes’ but these were lumped entities (for example, protein synthesis capacity) to reduce model complexity.
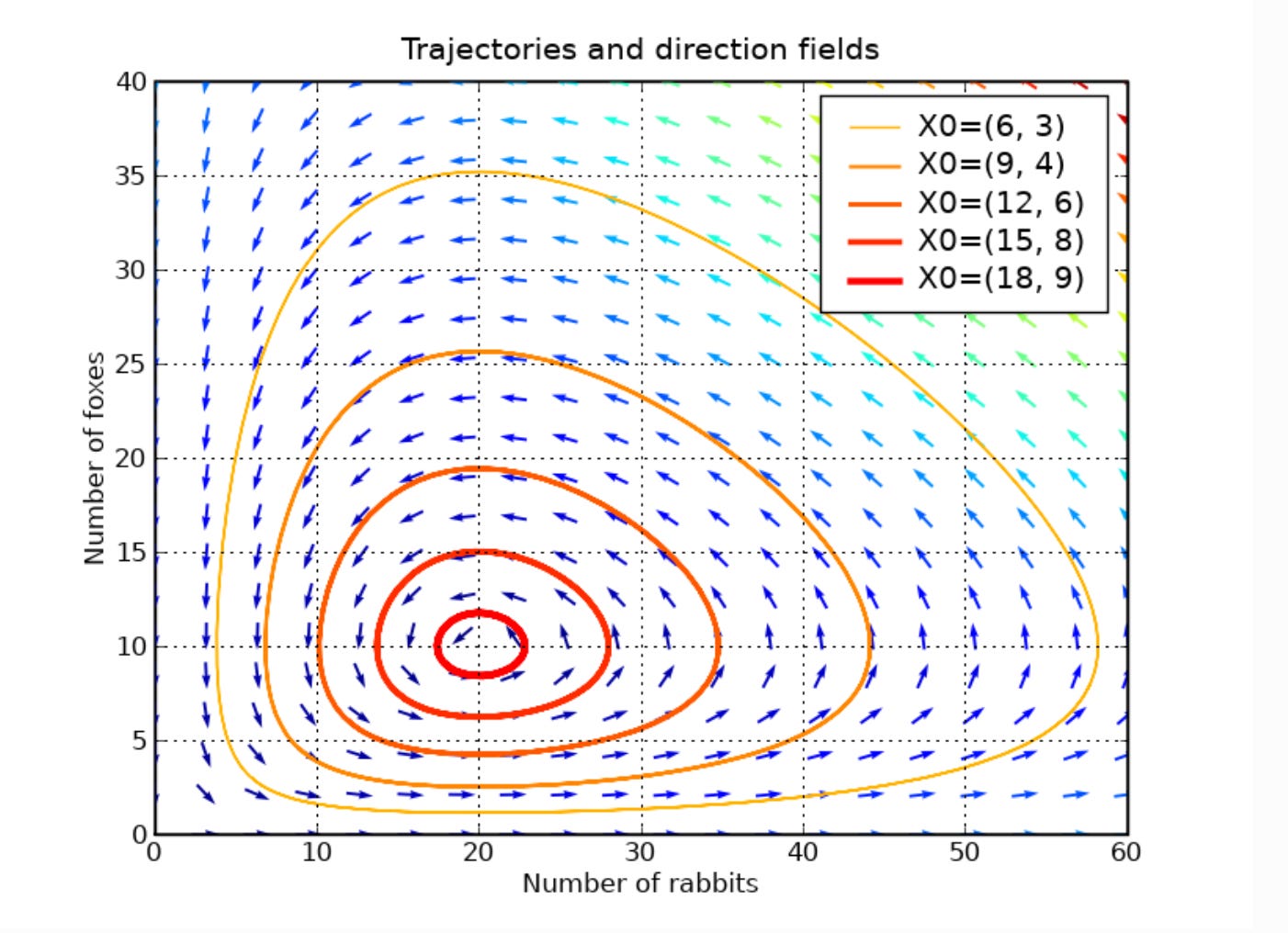
Hyun-Seob Song led the generation of a review article that covered all of the frameworks used to model microbial communities. Although 10 years old, it remains a good overview of the discrete approaches.
Where are we?
I find myself disappointed that mathematical modelling of microbial populations or communities has not advanced further. I found only 65 publications in what I consider high-quality journals over the past 5 years. Some of the reasons go back to my science-of-microbial-ecology-science post: current funding mechanisms do not particularly value this type of research. I also view the genome-scale flux-balance approach an intellectual dead end for microbial ecology – but almost half of those 65 publications used this.
Progress requires effective intellectual engagement between microbial ecologists and mathematicians. It would be rare for a single individual to function at the highest level in both ecology and mathematics. I go back to my collaborations with Ramki and his students at Purdue. Yes, the topic was a simpler one – modeling the activity and growth of a microbe in a bioreactor. We would sit and discuss the principles of microbial physiology relevant to the model and the mathematics behind implementing it. The students were very receptive to learning more about microbial physiology and I got to the point where I could read a page of ordinary differential equations and glean what they expressed. This required serious work on both sides to understand where they were coming from.
I had mentioned Hyun-Seob Song’s review of modeling of microbial communities earlier. He is a former colleague who exemplifies the type of traits necessary for successful collaborations in this area. He has a deep understanding of mathematics and a willingness to work together with microbiologists, biogeochemists and environmental scientists to advance this area. He is now a faculty member at the University of Nebraska-Lincoln, and I find the directions he is pursuing with colleagues of great interest.
A premise I developed in earlier posts was that microbial ecosystems are complex systems™️, in which dynamics are nonlinear and driven both by external perturbations (regular or stochastic) and internal (biological) dynamics. If that is so, it feels essential to make the mathematics of nonlinear dynamics and concepts such as limit cycles, attractors, bifurcations and regime shifts an essential part of modeling efforts — see Fujita et al. (2023) for a recent example.
References
Cockx BJR et al. (2024) Is it selfish to be filamentous in biofilms? Individual-based modeling links microbial growth strategies with morphology using the new and modular iDynoMiCS 2.0. PLoS Comput Biol 20(2): e1011303. https://doi.org/10.1371/journal.pcbi.1011303
Fujita H et al. (2023) Alternative stable states, nonlinear behavior,and predictability of microbiome dynamics. Microbiome 11:63. https://doi.org/10.1186/s40168-023-01474-5
May, RM (2001, revised) Stability and Complexity in Model Ecosystems. 301 pp. Princeton University Press;
Odum, HT. (1994) Ecological and general systems : an introduction to systems ecology. 644 pages. University Press of Colorado, Niwot, Colo., ©1994
Song H-S et al. (2014) Mathematical Modeling of Microbial Community Dynamics: A Methodological Review. Processes 2:711-752. doi:10.3390/pr2040711
Sugihara, G., & May, R. M. (1990). Nonlinear forecasting as a way of distinguishing chaos from measurement error in time series. Nature, 344: 734–741. https://doi.org/10.1038/344734a0
Some general overviews on modeling in microbial ecology
Kreft JU et al. (2017) From Genes to Ecosystems in Microbiology: Modeling Approaches and the Importance of Individuality. Front. Microbiol. 8:2299. doi: 10.3389/fmicb.2017.02299
Martinez-Rabert E et al. 2023 Multiscale models driving hypothesis and theory-based research in microbial ecology. Interface Focus 13: 20230008. https://doi.org/10.1098/rsfs.2023.0008
Vallina SM et al. (2019) Models in Microbial Ecology. Encyclopedia of Microbiology 4th edition, pp.211-246. Elsevier Inc.
van den Berg, N.I. et al. Ecological modelling approaches for predicting emergent properties in microbial communities. Nat Ecol Evol 6, 855–865 (2022). https://doi.org/10.1038/s41559-022-01746-7
Widder, S. et al. (2016) Challenges in microbial ecology: building predictive understanding of community function and dynamics. ISME J 10, 2557–2568. https://doi.org/10.1038/ismej.2016.45
Your moment of Zen
My procrastination in producing this post was a combination of being away for 3 weeks in Patagonia and recovering from that expedition. Patagonia is a fascinating area – as someone who lived in and visited low population density areas of Western North America, it was fascinating to travel ~4000 km by ship and see very few towns. Well, there were some occasional hot spots with creatures new to me — in this instance Emperor Cormorants